| Name: | Mouse monoclonal antibody to S. aureus CAS9 |
| Immunogen: | C-terminal region of S. aureus, amino acids 803-1053 of sequence CCK74173, expressed in and purified from E. coli. |
| HGNC Name: | N.A. |
| UniProt: | J7RUA5 |
| Molecular Weight: | 124kDa |
| Host: | Mouse |
| Isotype: | IgG1 heavy, κ light |
| Species Cross-Reactivity: | Staphylococcus aureus |
| RRID: | AB_2572247 |
| Format: | Purified antibody at 1mg/mL in 50% PBS, 50% glycerol plus 5mM NaN3 |
| Applications: | WB, IF/ICC, IHC |
| Recommended Dilutions: | WB: 1:1,000, IF/ICC: 1:1,000-1:5,000 |
| Storage: | Store at 4°C for short term, for longer term at -20°C |
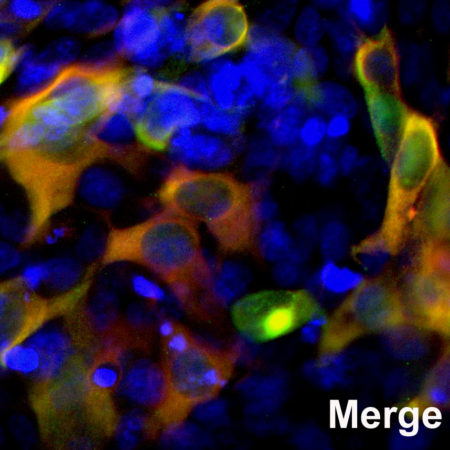
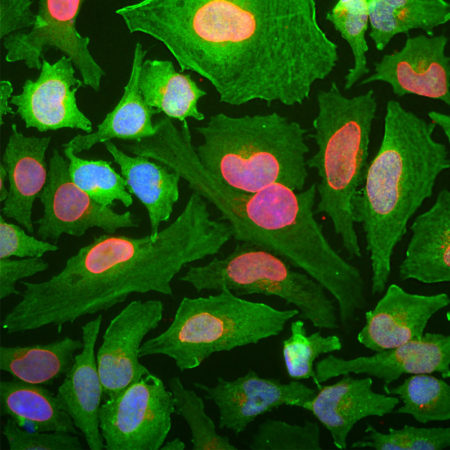
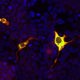
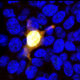
HEK293 cells were transfected with a fusion construct of green fluorescent protein (GFP) fused with the C-terminal sequence 803-1053 of S. aureus Cas9. Cells were stained with mouse mAb to S. aureusCas9, MCA-6F7, in red and the expressed GFP gives a green signal. The two signals overlap as expected. The nuclei of both transfected and untransfected cells are revealed by the DAPI DNA stain.
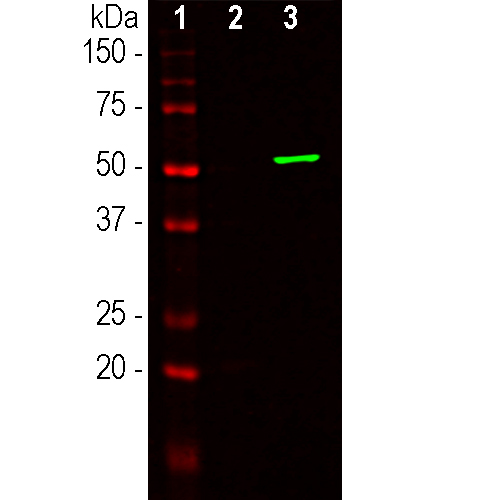
Western blot analysis of HEK293 cell lysates using mouse mAb to S. aureus CAS9, MCA-6F7: [1] protein standard (red), [2] non-transfected cells, and [3] transfected cells with GFP-Cas9 (C-terminal 803-1053 amino acids of S. aureus CAS9) fusion conctruct. Strong band at about 53kDa corresponds to the GFP-CAS9 fusion protein.
Mouse Monoclonal Antibody to S. aureus CAS9
Cat# MCA-6F7
$120.00 – $800.00
A recent revolution in biology has been stimulated by the discovery of CRISPR, or “Clustered Regularly Interspaced Short Palindromic Repeats” and the understanding of the “CRISPR Associated” enzymes (CAS 1,2). The CRISPR repeated sequences are found in bacterial genomes and function as part of unique bacterial immune system which contain short DNA sequences derived from viruses which have infected the bacteria. These virally derived sequences can make short RNA sequences which can hybridize with specific viral DNA and target a nuclease, such as CAS9, to the viral sequence. So CAS9 is directed to cleave the specific viral sequence and so inactivate the virus. The RNA sequence can be designed to specifically cut DNA virtually anywhere, including in the genomes of living human and other mammalian cells, allowing inexpensive gene editing with unprecedented ease. For example three groups of researchers essentially cured the disease state in a mouse model of Duchenne muscular dystrophy (3-5). A similar approach essentially cured dogs affected with a related disease state (6). Several varieties of CAS9 have been studied and there are several other related enzymes with similar properties. Much of the early work was performed with CAS9 from Streptococcus pyogenes which is rather large at ~158kDa, so the corresponding DNA is also rather large at about 4.2kb. This is problematic with some expression systems especially since DNA encoding RNA sequences and possibly other regulatory elements are usually required. The CAS9 gene of Staphylococcus aureus is significantly smaller, 3kb, producing a protein of 124kDa (7). For an excellent recent review of the various CAS family enzymes and their utility see reference 8.
The MCA-6F7 antibody was raised against the C-terminal 250 amino acids of S. aureus CAS9 in the sequence CCK74173. It can be used to verify expression of S. aureus CAS9 in cells and tissues. The antibody does not bind S. pyogenes CAS9. We used the same S. aureus immunogen to generate rabbit and chicken polyclonals to S. aureus CAS9, RPCA-CAS9-SA and CPCA-CAS9-SA. Mouse select image at left for larger view.
1. Hsu PD, Lander ES, Zhang F. Development and Applications of CRISPR-Cas9 for Genome Engineering. Cell 157:1262-78 (2014).
2. Doudna1 JA, Charpentier E. The new frontier of genome engineering with CRISPR-Cas9 Science 346:1077-86 (2014)
3. Long C, et al. Postnatal genome editing partially restores dystrophin expression in a mouse model of muscular dystrophy. Science 351:400-3 (2015).
4. Nelson CE, et al. In vivo genome editing improves muscle function in a mouse model of Duchenne muscular dystrophy. Science 351:403-7 (2015).
5. Tabebordbar M, et al. In vivo gene editing in dystrophic mouse muscle and muscle stem cells. Science 351:407-11 (2015).
6. Amoasii L. et al. Gene editing restores dystrophin expression in a canine model of Duchenne muscular dystrophy.
Science doi:10.1126/science.aau1549 (2018).
7. Ran FA, et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 520:186-91 (2015).
8. Knott GJ, Doudna J. CRISPR-Cas guides the future of genetic engineering. Science 361:866-9 (2018).
Related products
Contact info
EnCor Biotechnology Inc.
4949 SW 41st Boulevard, Ste 40
Gainesville
Florida 32608 USA
Phone: (352) 372 7022
Fax: (352) 372 7066
E-mail: admin@encorbio.com